AIxCell – Democratizing Deep Learning for Biomedical Image Analysis
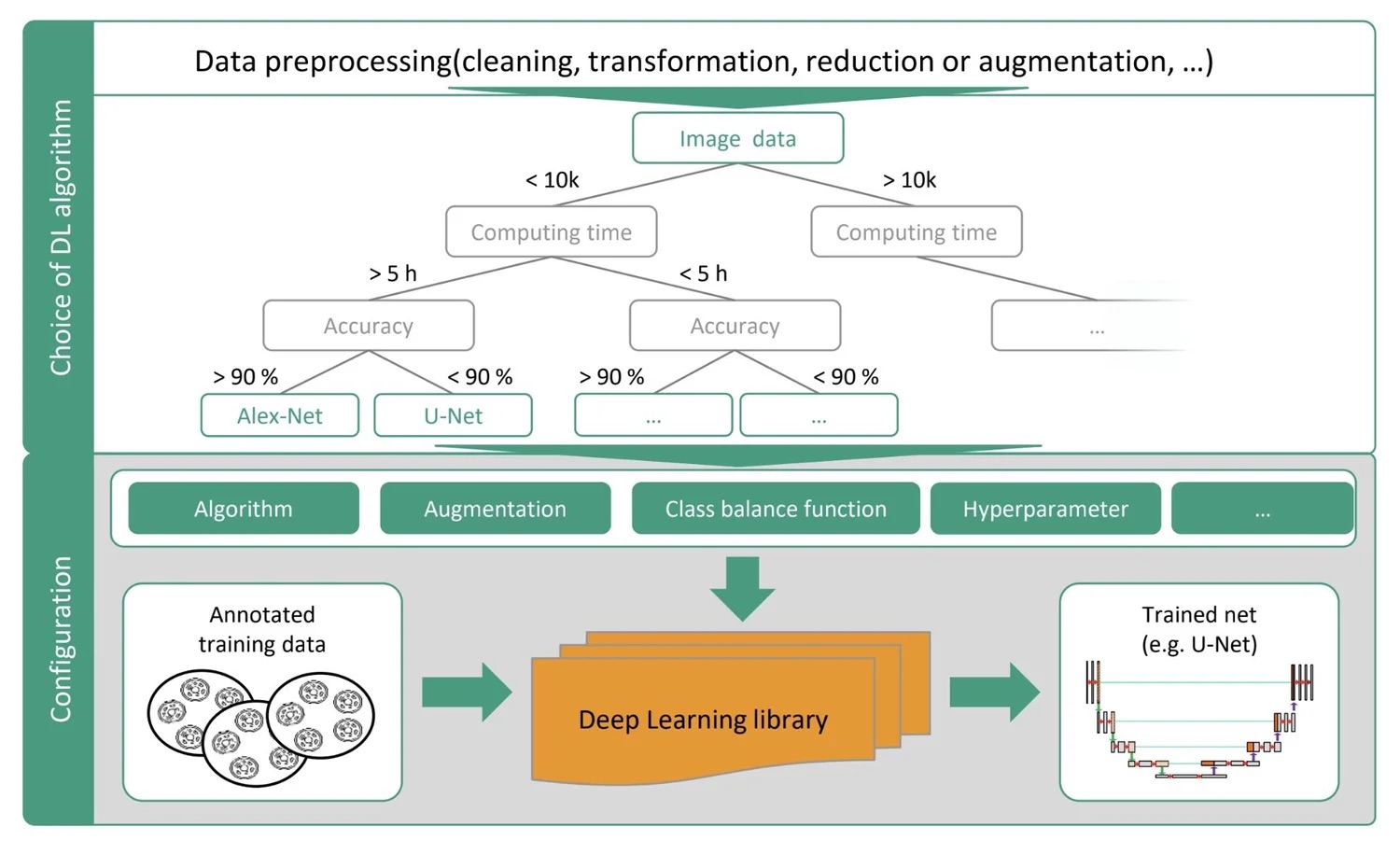
As part of the AIxCell research project, funded by the German Federal Ministry for Economic Affairs and Climate Action, I contributed to building an AutoML platform that empowers biomedical professionals to independently design, train, and deploy deep learning models for cell segmentation and classification without writing a single line of code. The platform's meta-learning engine automatically selects optimal algorithms based on task, data, and available resources, drastically reducing the time, cost, and expertise typically required for AI adoption in life sciences.
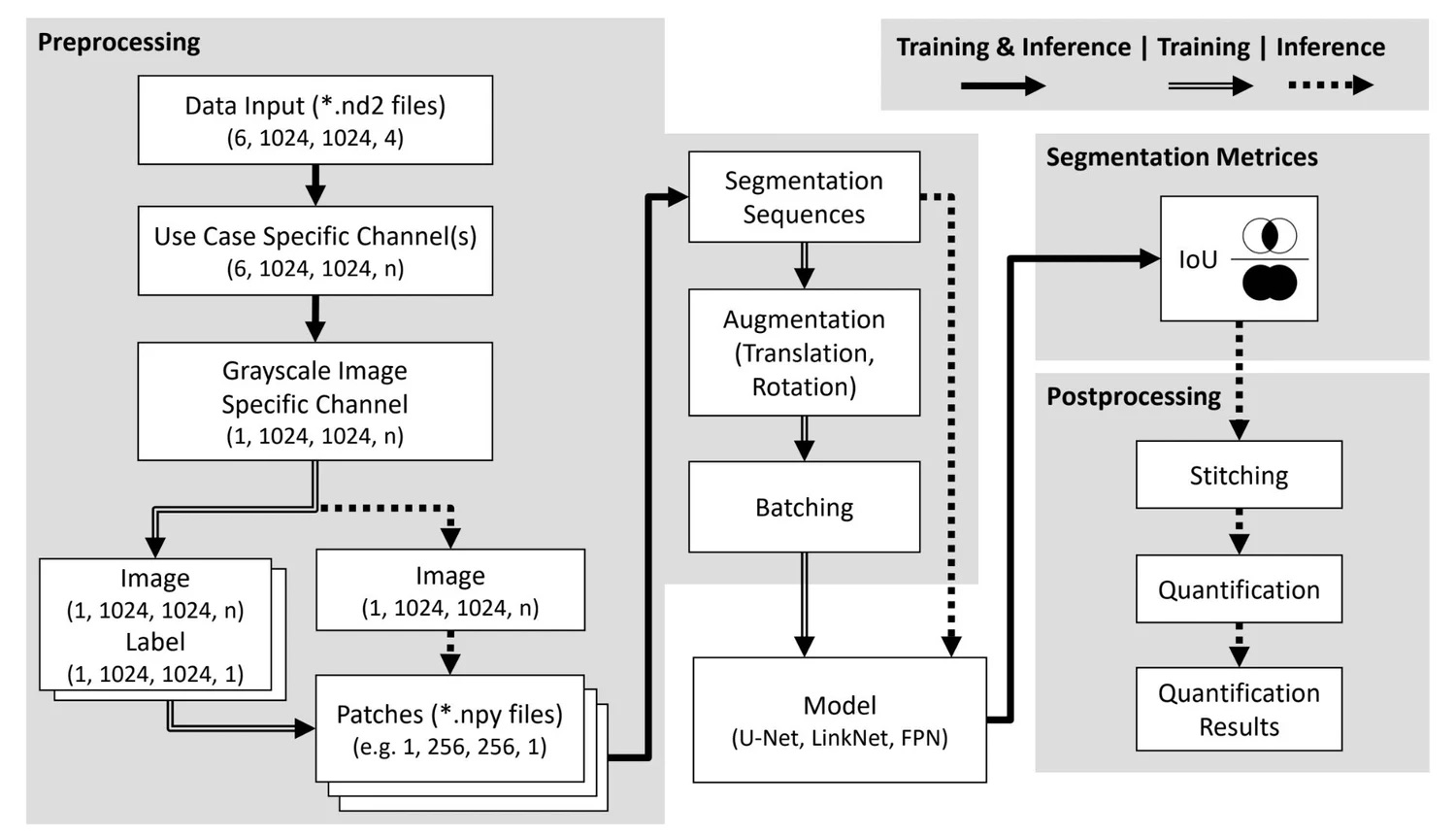
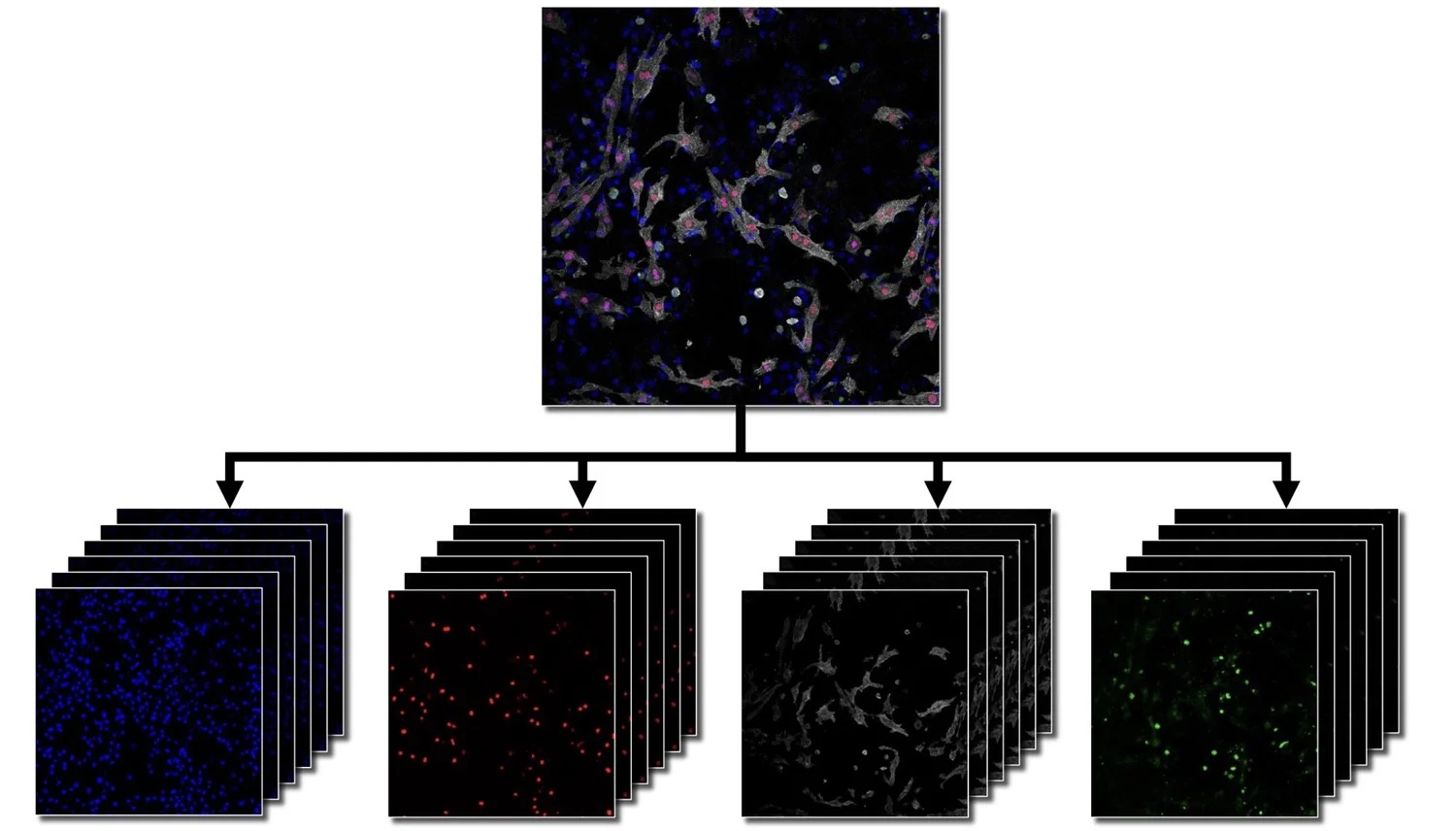
My contribution centered on my master's thesis: "Benchmarking of Deep Learning Algorithms for Stem Cell Classification". I developed a three-stage image analysis pipeline for cardiomyocyte segmentation in confocal microscopy, ran 173 experiments benchmarking 18 encoder-decoder architectures, and achieved test accuracies of up to 82% on small, high-complexity biomedical datasets. This work was published at MIDL 2022 in Zürich and directly enhanced AIxCell's model library and meta-learning capabilities.
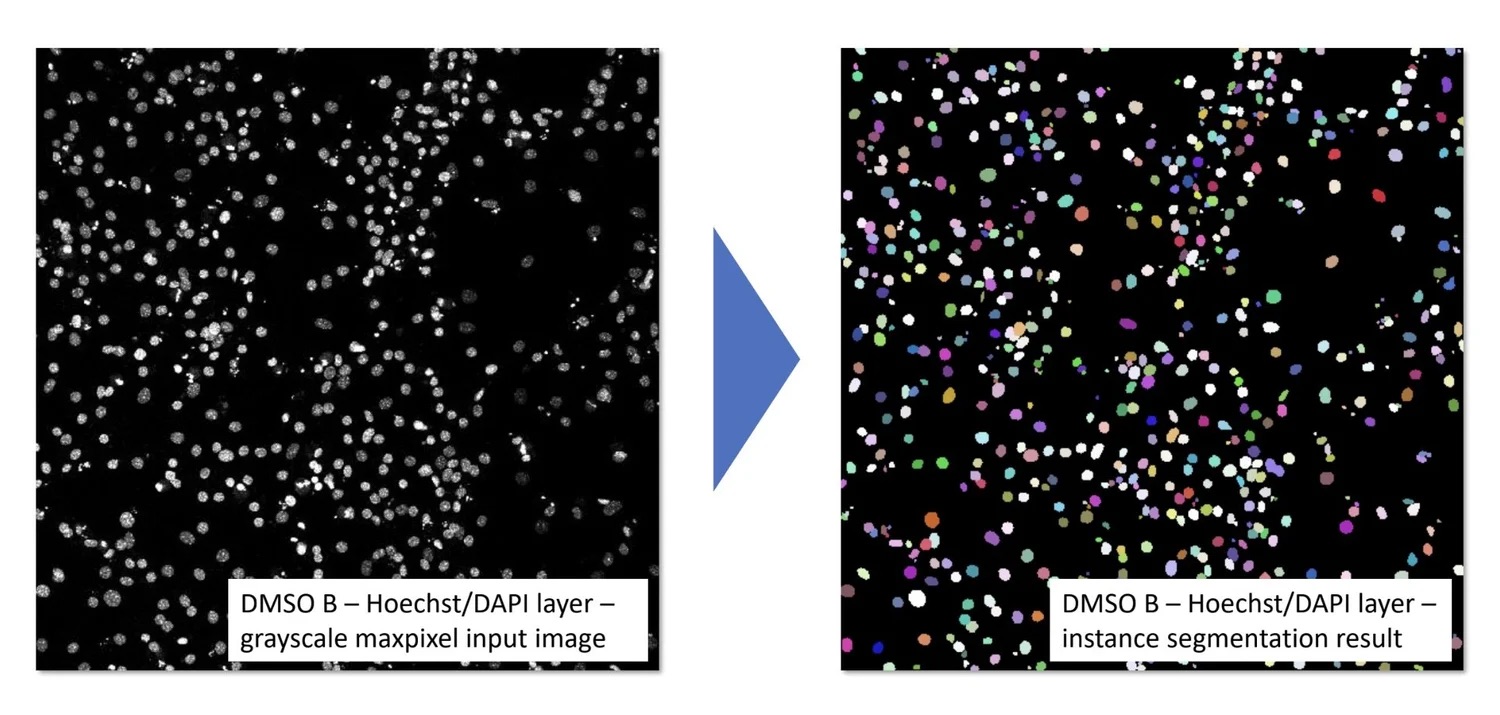

I contributed to this project as part of my master's thesis at Fraunhofer IPT, which led to a peer-reviewed publication at MIDL 2022.